missing translation for 'onlineSavingsMsg'
Learn More
Learn More
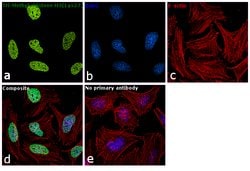
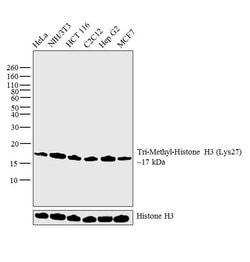
Invitrogen™ H3K27me3 Monoclonal Antibody (G.299.10), ChIP-Verified


Description
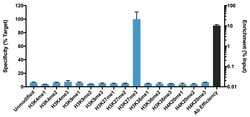
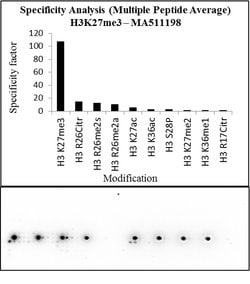
It is not recommended to aliquot this antibody. This antibody is not cross-reactive with mono-methylated, di-methylated or tri-methylated histone H3 at Lys4, Lys9, Lys36 or Histone H4 at Lys20. Click here for Master Lot linking information.
H3K27me3 is the epigenetic mark where histone H3 is trimethylated at lysine residue 27. H3K27me3 is an epigenetic silencing mark and thus associated with downregulation of nearby genes and often found in heterochromatin regions. H3K27me3 is mainly found at inactive gene promoters and generally is found in opposition to the activating mark H3K4me3. H3K27me3 often occupies gene rich regions with close association to development regulators. EZH2, a component of the PRC2 complex, is the histone methyltransferase (or writer) responsible for trimethylating H3K27. UTX and JMJD3 both are histone demethylases (or erasers) for H3K27me3. The regulation of H3K27me3 is crucial for gene regulation and dysregulation of H3K27me3 has been implicated in numerous diseases including cancer. H3K27me3 antibodies are often used for western blot and ChIP applications to study changes in levels and occupancy of H3K27me3.

Specifications
Specifications
| Antigen | H3K27me3 |
| Applications | Peptide Array, ChIP Assay, ChIP sequencing (ChIP-seq), Immunohistochemistry (Paraffin), Western Blot, Immunocytochemistry |
| Classification | Monoclonal |
| Clone | G.299.10 |
| Concentration | 102 μg/mL |
| Conjugate | Unconjugated |
| Formulation | 0.01M HEPES with 0.15M NaCl, 50% glycerol, 100μg/mL BSA and <0.02% sodium azide; pH 7.5 |
| Gene | H3C15 |
| Gene Accession No. | P68431, P68433, P84228, Q71DI3 |
| Gene Alias | H3; H3 histone; H3 histone family, member A; H3 histone family, member I; H3 histone family, member K; H3 histone family, member L; H3 histone family, member M; H3 histone, family 2; H3 histone, family 3A; H3 histone, family 3B; H3 histone, family 3B (H3.3B); H3 histone, family 3B.1; H3 histone, family 3C; H3 K27; H3.1-221; H3.1-291; H3.1-I; H3.2; H3.2-221; H3.2-614; H3.2-615; H3.2-616; h3.2a; H3.3A; H3.3b; H3.5; H3/A; H3/b; H3/d; H3/f; H3/i; H3/j; H3/k; H3/l; H3/M; H3/n; H3/o; H3-143; H3-291; H3-3A; H3-3b; h3-5; H3-53; H3-614; H3a; H3b; H3-B; H3C1; H3C10; H3c11; H3C12; H3C2; H3C3; H3C4; H3C6; H3C7; H3c8; H3f; H3-F; H3F1K; H3F2; H3F3; h3f3a; H3f3b; h3f3b.1; H3F3C; h3f3d; H3FA; H3FB; H3FC HIST1H3C; H3FD; H3FF; H3FH; H3FI; H3FJ; H3FK; H3FL; H3FM; H3FN; H3g; H3h; H3i; H3k27; H3Lys27me3; h3r; Hist1; Hist1h3a; HIST1H3B; Hist1h3c; Hist1h3d; HIST1H3E; HIST1H3F; hist1h3g; hist1h3g.L; HIST1H3H; Hist1h3i; HIST1H3J; hist2h3; HIST2H3A; Hist2h3b; HIST2H3C; Hist2h3c1; Hist2h3c2; Hist2h3c2-ps; Hist2h3ca1; Hist2h3ca2; HIST2H3D; histone 1, H3a; histone 1, H3b; histone 1, H3f; histone 1, H3h; histone 2, H3a; histone 2, H3c; histone 2, H3c2; histone 2, H3ca2; Histone 3; histone cluster 1, H3a; histone cluster 1, H3b; histone cluster 1, H3f; histone cluster 1, H3g protein L homeolog; histone cluster 1, H3h; histone cluster 2, H3a; histone cluster 2, H3c; histone cluster 2, H3c2; histone cluster 2, H3c2, pseudogene; histone gene complex 1; histone H3; histone H3.1; Histone H3.2; histone H3.3; Histone H3.3C; histone H3.5; Histone H3/a; Histone H3/b; Histone H3/c; Histone H3/d; Histone H3/f; Histone H3/h; Histone H3/i; Histone H3/j; Histone H3/k; histone H3/l; Histone H3/m; histone H3/o; Histone H3K27me3; histone variant H3.5; hypothetical protein LOC550262; M32461; methyl Histone 3; methyl Histone H3; PP781; Trimethyl H3; Tri-methyl H3; Trimethyl Histone H3; Tri-methyl Histone H3; Tri-Methyl-H3 K27; Tri-Methyl-H3 Lys27; Tri-Methyl-Histone H3 K27; Tri-Methyl-Histone H3 Lys27; wu:fa25h06; wu:fa96g06; wu:fb07a08; wu:fb36f01; XELAEV_18028537mg; zgc:110292; zgc:174300; zgc:56193; zgc:56418; zgc:64222; zgc:86731 |
| Show More |
Product Title
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.
Spot an opportunity for improvement?