missing translation for 'onlineSavingsMsg'
Learn More
Learn More
Invitrogen™ STARD13 Polyclonal Antibody


Description
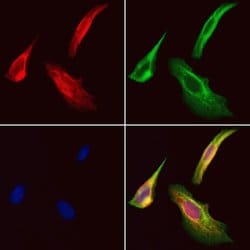
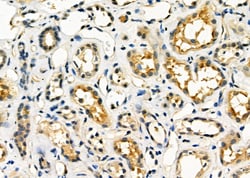
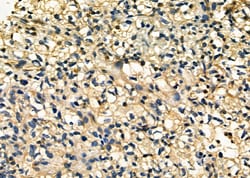
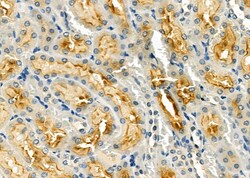
Antibody detects endogenous levels of total STA13.
This gene encodes a protein which contains an N-terminal sterile alpha motif (SAM) for protein-protein interactions, followed by an ATP/GTP-binding motif, a GTPase-activating protein (GAP) domain, and a C-terminal STAR-related lipid transfer (START) domain. It may be involved in regulation of cytoskeletal reorganization, cell proliferation, and cell motility, and acts as a tumor suppressor in hepatoma cells. The gene is located in a region of chromosome 13 that is associated with loss of heterozygosity in hepatocellular carcinomas. Alternatively spliced transcript variants encoding different isoforms have been described for this gene.

Specifications
Specifications
| Antigen | STARD13 |
| Applications | Immunohistochemistry (Paraffin), Western Blot, Immunocytochemistry |
| Classification | Polyclonal |
| Concentration | 1 mg/mL |
| Conjugate | Unconjugated |
| Formulation | PBS with 50% glycerol and 0.02% sodium azide; pH 7.4 |
| Gene | STARD13 |
| Gene Accession No. | Q923Q2, Q9Y3M8 |
| Gene Alias | 46H23.2; ARHGAP37; deleted in liver cancer 2 protein; DLC2; DLC-2; GT650; LINC00464; long intergenic non-protein coding RNA 464; Rho GTPase activating protein on chromosome 13q12; Rho GTPase-activating protein; serologically defined colon cancer antigen 13; serologically defined colon cancer antigen 28; STA13; StAR related lipid transfer domain containing 13; STARD13; StAR-related lipid transfer (START) domain containing 13; StAR-related lipid transfer domain containing 13; stAR-related lipid transfer protein 13; START domain-containing protein 13 |
| Gene Symbols | STARD13 |
| Show More |
Product Title
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.
Spot an opportunity for improvement?