missing translation for 'onlineSavingsMsg'
Learn More
Learn More
Invitrogen™ PITPNM1 Polyclonal Antibody


Description
Immunogen sequence: SRRGSMNNELL SPEFGPVRDP LADGVEGLGR GSPEPSALPP QRIPSDMASP EPEGSQNSLQ AA Highest antigen sequence identity to the following orthologs - mouse 83%, rat 83%.
Regulates RHOA activity, and plays a role in cytoskeleton remodeling. Necessary for normal completion of cytokinesis. Plays a role in maintaining normal diacylglycerol levels in the Golgi apparatus. Binds phosphatidyl inositol phosphates (in vitro). May catalyze the transfer of phosphatidylinositol and phosphatidylcholine between membranes. Necessary for maintaining the normal structure of the endoplasmic reticulum and the Golgi apparatus. Required for protein export from the endoplasmic reticulum and the Golgi. Binds calcium ions.

Specifications
Specifications
| Antigen | PITPNM1 |
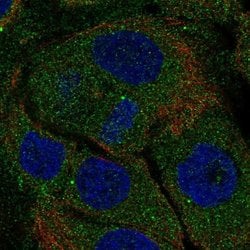
| Applications | Immunocytochemistry |
| Classification | Polyclonal |
| Concentration | 0.2 mg/mL |
| Conjugate | Unconjugated |
| Formulation | PBS with 40% glycerol and 0.02% sodium azide; pH 7.2 |
| Gene | PITPNM1 |
| Gene Accession No. | O00562 |
| Gene Alias | Dres9; drosophila retinal degeneration B homolog; Drosophila retinal degeneration B homolog 1; FLJ44997; membrane-associated phosphatidylinositol transfer protein 1; Mpt1; Mpt-1; NIR2; Nir-2; phosphatidylinositol membrane-associated; phosphatidylinositol transfer protein membrane associated 1; phosphatidylinositol transfer protein, membrane associated 1; phosphatidylinositol transfer protein, membrane-associated 1; Pitpnm; PITPnm 1; PITPNM1; Pyk2 N-terminal domain-interacting receptor 2; R75447; Rd9; RdgB; RDGB1; RDGBA; RDGBA1; retinal degeneration 9; retinal degeneration B alpha 1 |
| Gene Symbols | PITPNM1 |
| Show More |
Product Title
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.
Spot an opportunity for improvement?